I am doing a 3 way comparison of repeat elements in 3 species of anole lizards. All 3 of these genomes are Illumina Next-Gen. One challenge I face is moving the data around and formatting it properly. I had to do some Googling to find the appropriate bash commands, to process the fasta files from the terminal command line.
Below I:
1) cat - concatenated all the files into 1 large file
2) I want to sort by line size, but if you do it with a normal fasta file, the header is read as a different line than the sequence body (so it'll delete all the sequences under 250 INCLUDING the fasta headers, which have all the information I need!) Therefore, I had to remove all newlines \n and put in a * so the header and sequence all are read as one long line.
3) Next I used awk to sort based on size and I removed all sequences that are less than 250 base pairs.
4) Put the fasta format back the way it was before I sorted based on size.
#concatenate all the files from all 3 species into one sub-subfamily specific file
cat species1.fasta species2.fasta species3.fasta > /Desktop/masters_repeats/anolis_repeatfamily_3way_total.fasta
#i have to make all the entries one line, so i replaced the newline with a *
tr "()\n" "()*" < /Desktop/masters_repeats/anolis_repeatfamily_3way_total.fasta > /Desktop/masters_repeats/anolis_repeatfamily_3way_total_oneline.txt
sed -e 's/*>/\'$'\n>/g' /Desktop/masters_repeats/anolis_repeatfamily_3way_total_oneline.txt > /Desktop/masters_repeats/anolis_repeatfamily_3way_total_oneline2.txt
#sort based on size
awk '{ print length(), $0 | "sort -n" }' /Desktop/masters_repeats/anolis_repeatfamily_3way_total_oneline2.txt > /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength.txt
#i then removed all the lines that are under 250 characters
awk 'length > 250' /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength.txt > /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength_over250.txt
#remove everything before the >, which is just a space (because awk sorting adds an unecessary number to the beginning of each line. There may be a less verbose option, but I couldn't find it.)
sed 's/.* //' /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength_over250.txt > /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength_over250_clean.txt
#I replace the star with a newline again, so the fasta header, signified by a > is one line with the sequence below that.
tr "*" "\n" < /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength_over250_clean.txt > /Desktop/masters_repeats/anolis_repeatfamily_3way_total_SORTEDlength_over250_clean_expandedw-headers.fasta
My final product is ready to be uploaded into Geneious for alignments and tree-building.
Thursday, September 26, 2013
Wednesday, May 1, 2013
Primer Design Cont.
I am continuing to design probes for In Situ
Hybridization. Currently, I used Ensembl and BLAST to find gene
orthologues in the Tiger Salamander (Ambystoma). See Making Primers Part 1
for reference. After I received the primers, I tested them out via a
PCR reaction and my cDNA template (generated kindly by our undergrad
Minami Tokuyama). 7 out of 10 of the primers seem to work well. This is
continuation of the blog Primer Testing and Running a Gel. Note that this article is not peer reviewed and is subject to change.
Now that I have my gel, I can begin to analyze the results. 3 of the expected bands do not show up. The other 7 look good, as there are no strong multiple bands that would indicate non-specific binding. The 100 bp ladder can be read from the bottom up, with the first band in the first well as 100 bp, then next as 200 bp and so on.
Recall that small sequences move more quickly during gel electrophoresis, and should travel much farther from their starting point (the top of the gel) towards the bottom. Large sequences are slow and won't move as far. Therefore, the 5th band from the bottom of 100 bp DNA ladder, which is much brighter, indicates PCR products that are roughly 500 base pairs. I can see that seven of my gene products are between 400 and 600 basepairs.
This is good news to me, as this was the expected outcome.
I consult my spreadsheet to see what the predicted PCR product sizes are. I had previously determined this information by entering my primer sequences (forward and reverse) into the Sequence Manipulation suite: PCR Test, at bioinformatics.org.
I can see that in well 2, which is pax6 (exon2, 2nd primer set), should be 404 base pairs. I can see that in well 4, mef2a (exon1, 2nd primer set) should be the largest at 546 base pairs. Indeed, it is the largest sequence and has moved the slowest (from top to bottom, or from the black negative anode to red positive cathode). Most of the products should be around roughly 450 bp and this is what my gel reflects.
I can see faint Primer Dimer bands which are located under the 100 bp band (towards the bottom). This doesn't prevent me from using the primers; however, bands over 100 bp are troublesome as they might indicate the primers are binding nonspecifically.
After the primers have been tested and deemed correct, based on predicted PCR product size, reorder the primer with T7.
Now that I have my gel, I can begin to analyze the results. 3 of the expected bands do not show up. The other 7 look good, as there are no strong multiple bands that would indicate non-specific binding. The 100 bp ladder can be read from the bottom up, with the first band in the first well as 100 bp, then next as 200 bp and so on.
Recall that small sequences move more quickly during gel electrophoresis, and should travel much farther from their starting point (the top of the gel) towards the bottom. Large sequences are slow and won't move as far. Therefore, the 5th band from the bottom of 100 bp DNA ladder, which is much brighter, indicates PCR products that are roughly 500 base pairs. I can see that seven of my gene products are between 400 and 600 basepairs.
This is good news to me, as this was the expected outcome.
I consult my spreadsheet to see what the predicted PCR product sizes are. I had previously determined this information by entering my primer sequences (forward and reverse) into the Sequence Manipulation suite: PCR Test, at bioinformatics.org.
I can see that in well 2, which is pax6 (exon2, 2nd primer set), should be 404 base pairs. I can see that in well 4, mef2a (exon1, 2nd primer set) should be the largest at 546 base pairs. Indeed, it is the largest sequence and has moved the slowest (from top to bottom, or from the black negative anode to red positive cathode). Most of the products should be around roughly 450 bp and this is what my gel reflects.
I can see faint Primer Dimer bands which are located under the 100 bp band (towards the bottom). This doesn't prevent me from using the primers; however, bands over 100 bp are troublesome as they might indicate the primers are binding nonspecifically.
After the primers have been tested and deemed correct, based on predicted PCR product size, reorder the primer with T7.
Primer Testing and Running a Gel
MAKING STOCKS
The primers will come dry, in small tubes. They are stable at room temperature. As soon as you add water, you will want to keep the primers on ice. You will need to make a 200 millimollar stock of the primers. I also recommend making a working stock as well.
To make the 200 mm stock, find the volume in nanomoles and multiply by 5.
For example:
Gene1-1F - 29.4 nm x 5 = 147 ul
Gene1-1R - 35.1 nm x 5 = 175.5 ul
Gene2-1F - 29.6 nm x 5 = 148 ul
Gene2-1R - 29.2 nm x 5 = 146 ul
Take that amount of microliters per tube and add Molecular Grade ddH20 to the tube. You now have a 200 mm stock.
Vortex the tube to mix them. Spin them down in the centrifuge.
Working Stock
For regular PCR, the concentration should be at 20 mm. For RT-PCR, you want it to be at 10 mm.
Using the C1V1=C2V2 equation : (200 mM) x = (20 mM) (100 ul)
x = 10 ul
You want a working stock of 100 microliters. You can put both the forward and reverse primer into your working stock. So 10 ul/2 = 5 ul of the forward primer and 5 ul of the reverse primer.
Add 90 ul of Molecular grade water
5 ul of forward primer
5 ul of reverse primer
__________________________
Total of 100 ul working stock
Vortex the tube to mix them. Spin them down in the centrifuge.
You want to keep both the long-term 20 mm stock and the working stock (F+R primers) on ice, or in that -20 to -30 C freezer.
MASTERMIX
Next, you want to create a Master Mix solution. You want the number of reactions + 10% room for error. Total volume for each well (each primer being tested) is 20 ul. If you are testing 5 genes, then that would be 5 reactions (or 5 wells). So take the Mastermix recipe and times it by number of reactions and 1.1 for each of the elements.
Do not add the primer working stocks to the Mastermix. If you are using different templates, say cDNA from mouse and cDNA from frog. DO NOT ADD IN THE TEMPLATE yet to the Mastermix. However, if you are testing genes for the same species, with the same template you can add the template to the Mastermix. Let's assume that all the 5 genes I am testing are for the same species. I will add 22 ul of template to the Mastermix solution.
Next, get some tiny PCR tubes (and lids). Remember, the desired volume for each PCR tube is 20 ul (For regular PCR. For RT-PCR, you want 10 mm). You want to add 19.2 ul of Mastermix to each tube. Then you want to add 0.8 ul of each primer working stock (changing pipette tips each time!!). Then you total volume is 20 ul.
RUNNING THE PCR MACHINE
Each PCR machine varies, but here is the general procedure:
Put the tubes in. Tighten the lid.
-Tubes
-20 ul volume
-Heated lid
-Last step should run at 4C forever.
RUNNING A GEL
Right before you want to run the gel, add a loading dye to each well. I use a 6x loading dye, but it needs to be at 1x. I use the C1V1=C2V2 formula to solve for x:
(6x)(y) = (20 ul)(1x)
y = 3.333 ul
I want to add 3.3 ul of 6x loading dye to each tube. This will turn the solution from clear to blue. Spin the tubes down.
RUNNING A GEL
First you want to make the 1-2% agarose gel.
Depending on your gel casting tray size, you will need different volumes. I am running a small gel and my casting tray holds about 75 ml. To test how much volume it holds, just pour water in from a graduated cylinder to find out. I am going to make slightly more than I need to be safe. A clamp is used on the gel casting tray. Put the comb in. This will create wells when you pour the agarose in.
If I want 82 mls of agarose gel mix, I will use 0.82 grams of agarose powder.
I add the powder to 82 mls of Borax. Next I want to microwave it, until it boils. DON'T LET IT BOIL OVER! You may have to stop and wait a few times, until all the particles are dissolved. The solution below needs to be microwaved longer.
You want the solution to be clear.
When the solution is clear, wait for the agarose gel solution to cool down. When it is room temp, you want to add Ethidium.
Ethidium is a dangerous chemical and is usually kept in a safe place, stored at room temperature. For a small gel, use 1-2 ul of Ethidium. For a large gel, use 5 ul. In our lab, we use a pipetter reserved for the Ethidium only and kept in a cleared out area assumed to be ethidium-dirty.
Swirl the bottle to mix up the Ethidium into the agarose gel. Pour the gel in the gel casting tray, as pictured above. It should be 15 minutes or longer for the gel to congeal. If you wait too long it will solidify in the bottle.
MAKE SURE THAT NO BUBBLES ARE NEAR THE COMB!
When the gel is solid, you can unclamp the casting tray and carefully remove the tray with the agarose gel in it. It should be rectangular and jelly-like.
Move the gel to the gel electroporesis box.
Pour in Borax to cover the tray. Don't fill it past the MAX line.
Pull the comb out. If you place a dark sheet under the box, you should be able to see square wells. This is where you will pipette your mix into.
For a large gel, you want to put 8-10 ul of each Mastermix+Template+primers+loading dye into each well. For smaller gels, 2-4 ul might be adequate. If you run your gel and get blobs and trails, it may be because there is too much DNA. Cutting the concentration will help.
Use a different pipette tip each time!
In the peripheral wells, you want to use a DNA ladder. I use a 100 bp ladder and a 1000 bp ladder. This will allow you to gauge how large your product is compared to a reliable measure of size.
In general, you can use about 5 microliters for the 100 bp ladder and 2-3 microliters for the 1000 bp ladder.
Write down which primer is in each well!
Put the cover onto the electroporesis, matching black to black and red to red.
Turn the gel electroporesis machine on. I use a setting of 110, then start it.
You should see bubbles coming out of the anodes. Let your gel run for 20-45 minutes. You want them to run down 75% the length of your gel. If you let it run too long, it will run right off your gel!
A finished gel should look like the one below:
Note: I disposed of my gel in a special hazardous waste ethidium bromide container.
Next, I photograph the gel using a special Gel-Doc machine that uses Trans-UV to image. I invert the image when exporting as a .TIFF file to get the picture below:
INTERPRETING A GEL
Recall that bigger fragments move more slowly and smaller fragments move faster. To give you an idea of relative size, the DNA ladders (100 bp and 1000 bp) are used. Since the fragments will run from the negative electrode (black) to the positive electrode (red), the slower fragments will be closer to the starting point. Read the ladder from the bottom up. See below. The brighter bands are 500 and 1000.
You want your band to be a single, bold band. Multiple bands can indicate contamination or non-specific binding of your primers.

Attribution-ShareAlike CC BY-SA
This license lets others remix, tweak, and build upon your work even for commercial purposes, as long as they credit you and license their new creations under the identical terms. This license is often compared to “copyleft” free and open source software licenses. All new works based on yours will carry the same license, so any derivatives will also allow commercial use.
The primers will come dry, in small tubes. They are stable at room temperature. As soon as you add water, you will want to keep the primers on ice. You will need to make a 200 millimollar stock of the primers. I also recommend making a working stock as well.
To make the 200 mm stock, find the volume in nanomoles and multiply by 5.
 |
| Primer stock |
For example:
Gene1-1F - 29.4 nm x 5 = 147 ul
Gene1-1R - 35.1 nm x 5 = 175.5 ul
Gene2-1F - 29.6 nm x 5 = 148 ul
Gene2-1R - 29.2 nm x 5 = 146 ul
Take that amount of microliters per tube and add Molecular Grade ddH20 to the tube. You now have a 200 mm stock.
Vortex the tube to mix them. Spin them down in the centrifuge.
Working Stock
For regular PCR, the concentration should be at 20 mm. For RT-PCR, you want it to be at 10 mm.
Using the C1V1=C2V2 equation : (200 mM) x = (20 mM) (100 ul)
x = 10 ul
You want a working stock of 100 microliters. You can put both the forward and reverse primer into your working stock. So 10 ul/2 = 5 ul of the forward primer and 5 ul of the reverse primer.
Add 90 ul of Molecular grade water
5 ul of forward primer
5 ul of reverse primer
__________________________
Total of 100 ul working stock
Vortex the tube to mix them. Spin them down in the centrifuge.
You want to keep both the long-term 20 mm stock and the working stock (F+R primers) on ice, or in that -20 to -30 C freezer.
MASTERMIX
Next, you want to create a Master Mix solution. You want the number of reactions + 10% room for error. Total volume for each well (each primer being tested) is 20 ul. If you are testing 5 genes, then that would be 5 reactions (or 5 wells). So take the Mastermix recipe and times it by number of reactions and 1.1 for each of the elements.
Do not add the primer working stocks to the Mastermix. If you are using different templates, say cDNA from mouse and cDNA from frog. DO NOT ADD IN THE TEMPLATE yet to the Mastermix. However, if you are testing genes for the same species, with the same template you can add the template to the Mastermix. Let's assume that all the 5 genes I am testing are for the same species. I will add 22 ul of template to the Mastermix solution.
Next, get some tiny PCR tubes (and lids). Remember, the desired volume for each PCR tube is 20 ul (For regular PCR. For RT-PCR, you want 10 mm). You want to add 19.2 ul of Mastermix to each tube. Then you want to add 0.8 ul of each primer working stock (changing pipette tips each time!!). Then you total volume is 20 ul.
RUNNING THE PCR MACHINE
Each PCR machine varies, but here is the general procedure:
Put the tubes in. Tighten the lid.
-Tubes
-20 ul volume
-Heated lid
-Last step should run at 4C forever.
Check
with your lab about the specific program you want to run (with temps,
times, and cycles). Our PCR machine has a program pre-set.
Here is the program I use for PCR.
5 minutes 94 degrees (x 1 cycle)
1 minute 94 degrees (x 40 cycles)
1 minute 55 degrees (x 40 cycles)
2 minutes 72 degrees (x 40 cycles)
7 minutes 72 degrees (x 1 cycle)
15 minutes 4 degrees (x 1 cycle)
(set on 4 degrees forever, if you want to leave it overnight)
If you want to know what occurs during each step (of the denaturation, annealing, and extension process), check out NCBI's explanation.
RUNNING A GEL
Right before you want to run the gel, add a loading dye to each well. I use a 6x loading dye, but it needs to be at 1x. I use the C1V1=C2V2 formula to solve for x:
(6x)(y) = (20 ul)(1x)
y = 3.333 ul
I want to add 3.3 ul of 6x loading dye to each tube. This will turn the solution from clear to blue. Spin the tubes down.
 |
| Mastermix + Primer and Templates in PCR reaction tubes |
RUNNING A GEL
First you want to make the 1-2% agarose gel.
Depending on your gel casting tray size, you will need different volumes. I am running a small gel and my casting tray holds about 75 ml. To test how much volume it holds, just pour water in from a graduated cylinder to find out. I am going to make slightly more than I need to be safe. A clamp is used on the gel casting tray. Put the comb in. This will create wells when you pour the agarose in.
 |
| Gel Casting tray |
If I want 82 mls of agarose gel mix, I will use 0.82 grams of agarose powder.
I add the powder to 82 mls of Borax. Next I want to microwave it, until it boils. DON'T LET IT BOIL OVER! You may have to stop and wait a few times, until all the particles are dissolved. The solution below needs to be microwaved longer.
 |
| 1% Agarose Gel solution |
You want the solution to be clear.
When the solution is clear, wait for the agarose gel solution to cool down. When it is room temp, you want to add Ethidium.
Ethidium is a dangerous chemical and is usually kept in a safe place, stored at room temperature. For a small gel, use 1-2 ul of Ethidium. For a large gel, use 5 ul. In our lab, we use a pipetter reserved for the Ethidium only and kept in a cleared out area assumed to be ethidium-dirty.
Swirl the bottle to mix up the Ethidium into the agarose gel. Pour the gel in the gel casting tray, as pictured above. It should be 15 minutes or longer for the gel to congeal. If you wait too long it will solidify in the bottle.
MAKE SURE THAT NO BUBBLES ARE NEAR THE COMB!
When the gel is solid, you can unclamp the casting tray and carefully remove the tray with the agarose gel in it. It should be rectangular and jelly-like.
Move the gel to the gel electroporesis box.
Pour in Borax to cover the tray. Don't fill it past the MAX line.
Pull the comb out. If you place a dark sheet under the box, you should be able to see square wells. This is where you will pipette your mix into.
For a large gel, you want to put 8-10 ul of each Mastermix+Template+primers+loading dye into each well. For smaller gels, 2-4 ul might be adequate. If you run your gel and get blobs and trails, it may be because there is too much DNA. Cutting the concentration will help.
Use a different pipette tip each time!
In the peripheral wells, you want to use a DNA ladder. I use a 100 bp ladder and a 1000 bp ladder. This will allow you to gauge how large your product is compared to a reliable measure of size.
In general, you can use about 5 microliters for the 100 bp ladder and 2-3 microliters for the 1000 bp ladder.
 |
| Wells are filled in the gel electroporesis box |
Write down which primer is in each well!
Put the cover onto the electroporesis, matching black to black and red to red.
Turn the gel electroporesis machine on. I use a setting of 110, then start it.
You should see bubbles coming out of the anodes. Let your gel run for 20-45 minutes. You want them to run down 75% the length of your gel. If you let it run too long, it will run right off your gel!
A finished gel should look like the one below:
Note: I disposed of my gel in a special hazardous waste ethidium bromide container.
Next, I photograph the gel using a special Gel-Doc machine that uses Trans-UV to image. I invert the image when exporting as a .TIFF file to get the picture below:
INTERPRETING A GEL
 |
| Gel Doc |
Recall that bigger fragments move more slowly and smaller fragments move faster. To give you an idea of relative size, the DNA ladders (100 bp and 1000 bp) are used. Since the fragments will run from the negative electrode (black) to the positive electrode (red), the slower fragments will be closer to the starting point. Read the ladder from the bottom up. See below. The brighter bands are 500 and 1000.
You want your band to be a single, bold band. Multiple bands can indicate contamination or non-specific binding of your primers.
Attribution-ShareAlike CC BY-SA
This license lets others remix, tweak, and build upon your work even for commercial purposes, as long as they credit you and license their new creations under the identical terms. This license is often compared to “copyleft” free and open source software licenses. All new works based on yours will carry the same license, so any derivatives will also allow commercial use.
Thursday, January 17, 2013
Making Phylogenetic Tree Figures
Making phylogenetic trees takes many steps and requires the use of several online resources.
iTOL, or the Interactive Tree of Life, will automatically generate a tree of life based on NCBI identifiers. To use iTOL you will need the NCBI scientific name (including proper capitalization); just replace the space with an underscore. If you aren't sure about a species scientific name, you can search Ensembl, NBCI, or even Wikipedia.
Here is what I used to look at amniote evolution.
Mus_musculus
Homo_sapiens
Gallus_gallus
Taeniopygia_guttata
Alligator_mississippiensis
Xenopus_laevis
Trachemys_scripta
Pelodiscus_sinensis
Anolis_carolinensis
Danio_rerio
Homo_sapiens
Gallus_gallus
Taeniopygia_guttata
Alligator_mississippiensis
Xenopus_laevis
Trachemys_scripta
Pelodiscus_sinensis
Anolis_carolinensis
Danio_rerio
HOW DO I GENERATE THE TREE?
Enter in the scientific names and click generate tree. iTOL has many
features, which you can explore. I didn't particularly like their user
interface, so I simply used them to give me the Newick text. Newick text is just a way to represent trees in a language computers can easily 'read.'
After you generate the tree, you will be given Newick text, that establishes the tree structure. You can use the taxonomy IDS or scientific names. If the internal nodes are expanded, you will have a very large and detailed tree.
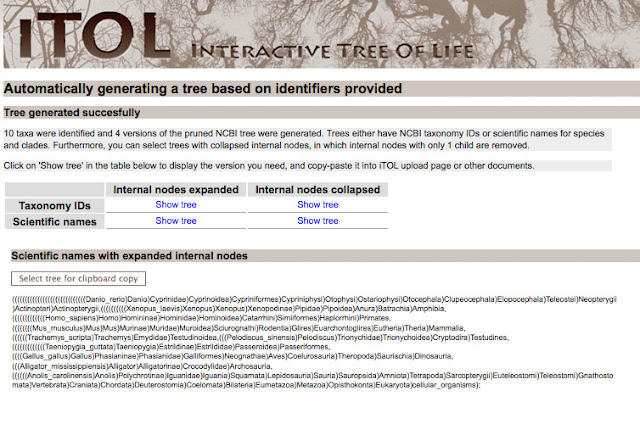
For my purposes, I wanted to a tree with the internal nodes collapsed. For making a publication quality figure, it's less crowded.
Next, I copied the text and pasted the following text into University of Indiana's Phlyodendron.
(((Xenopus_laevis,((Mus_musculus,Homo_sapiens) ,((Pelodiscus_sinensis,Trachemys_scripta) ,(Anolis_carolinensis,((Gallus_gallus,Taeniopygia_guttata) ,Alligator_mississippiensis)Arch)Sauria)Saurop)Amniota)Tetra,Danio_rerio) );
The original text was actually:
(((Xenopus_laevis,((Mus_musculus,Homo_sapiens)Euarchontoglires,((Pelodiscus_sinensis,Trachemys_scripta)Cryptodira,(Anolis_carolinensis,((Gallus_gallus,Taeniopygia_guttata)Neognathae,Alligator_mississippiensis)
Archosauria)Sauria)Sauropsida)Amniota)Tetrapoda,Danio_rerio)Euteleostomi);
But I replaced Euteleostomi, Neognathae, Cryptodira, Euarchontoglires with spaces. I also abbreviated a few names, so they didn't intersect with the lines. I want my final figure to look clean. Next, I chose to output a phenogram tree.
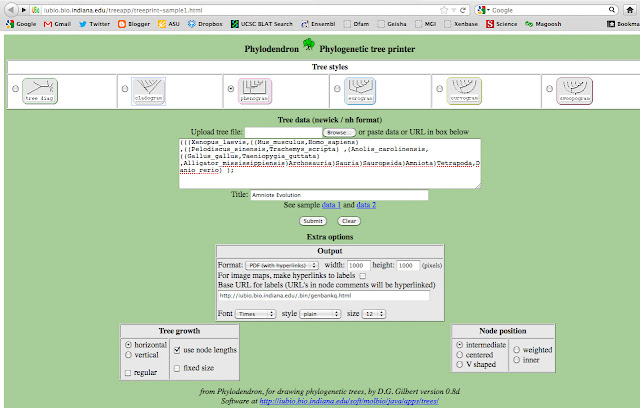
That will generate a PDF file, that looks like this:
HOW DO I MAKE IT LOOK GOOD?
Now, I use Photoshop to spruce up the image. First, I make the image I want to create a time line on the bottom and check the dates of each node. In general, I will compact the vertical lines to make it tighter. This is where a bit of awareness in image composition comes in handy. You don't want your image too look too spaced out or too crowded. Choose a color scheme that is not too jarring or too pale. The image composition should not distract from the information you are trying to convey!
USE LAYERS
When you begin adding features to your Photoshop file, you will want to make a new layer for each item, name it and keep track of what layer you are working on at all times. Keep a separate layer for the tree, the time scale, the boxes, each name, etc. You will thank me later! Save it as a .psd file.
SAVE MANY, SAVE OFTEN
I also like to save different versions every time I make a radical change. v_1, v_2, v_3. There are countless times I have had to use a backup file.
I recommend using Illustrator if you want a high-quality
publication-ready figure. The use of vectors in your images will allow
the image to still look great at different sizes. You can open your .psd Photoshop file in Illustrator. My general design was based on Sudhir Kumar's TimeTrees. It's a wonderful site, accompanied by a wonderful book and I highly recommend checking it out.
VECTORS ARE BETTER
If you use Adobe
Illustrator, you can save your image as a PDF file. Try zooming in and
out of your image. I am not going to give a lecture on what vectors are (Google it!), but with vectors, instead of bmps, the image will still
look great at many scales (instead of pixelated). This is especially true for the text, which often becomes distorted.
KNOW YOUR FILE SIZE REQUIREMENTS
If you are making the figure for a publication, you will need to consult their graphic or artwork guide. There are usually 3 sizes you can make your image:
1) Single column width in a double column paper
2) One and a half width
3) Full page width
For each of these sizes, you should make sure the image resolution is up to par. I like to have at least 300 resolution and a large document size. In general, if an image looks great when it's big, it will continue to look great as you shrink it down. The same is not true if you do it the other way around.
While each publication company has different requirements, some general sizes are:
DESIRED SIZE SIZE DPI
Single column width - 90 mm ~ 3500
One and half page width - 140 mm ~ 5500
Full page width - 190 mm ~ 7500
WHAT DATES SHOULD I USE?
There is, of course, always controversy in obtaining accurate evolutionary dates. They are estimates, at best, and having several sources to base your estimates off of is the best strategy. I love using the Time Tree program. It pulls in many sources indicating species molecular estimates. You can click on each paper and decide on what date you want to use for your tree.
Good luck with your tree making!
Monday, January 14, 2013
Making Primers, Pt I
Making primers is a long process. In Part I, I am just going to cover how to order the initial oligos.
If you are looking de novo for orthologue (gene equivalents) in another species, you may have to do some BLASTS to try to find them, including BLASTs for proteins, mRNA or highly conserved regions (like a promoter), depending on the amount of time diverged.
To begin, you have to search for the genes you want and save the sequence to a file. If you use Ensembl, you can search for a gene in a species and use the gene browser to visualize the gene structure. For example, I searched for Pax6 in the frog Xenopus. I can see right away that there are two isoforms of this gene.
So I know that the gene is spliced alternatively in two forms, which may have tissue specificity or functional importance. One isoform may be predominantly expressed, while another is found in low levels. Ideally, I want to capture them both. I will choose exons that are common to them both. For Pax6, the last 2 exons appear to be similiar enough. I want to export the exon sequence for two exons from Ensembl.
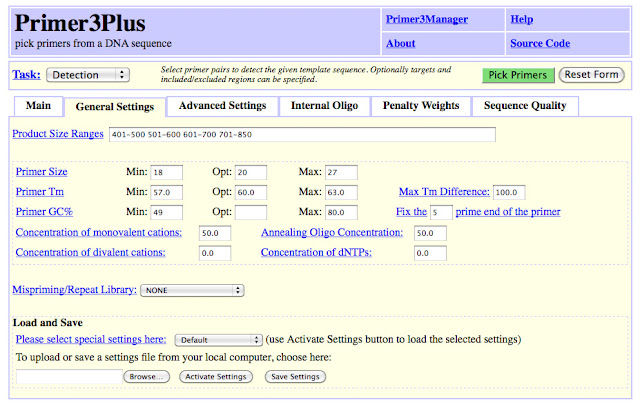
I want my oligo to span 2 exons ideally, such that the sequence spans an intron on either side. After each > is an exon. I find the 2 exons I want to use and copy/paste them into Primer3Plus.
I paste the exons into a text file and begin looking for a a good stretch, that spans introns, has a good GC content, and will give an appropriate product size.
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100940 exon1:KNOWN_protein_coding
ATGTCCCTAGGTCACAGCGGAGTCAATCAACTCGGGGGAGTGTTTGTGAACGGCCGACCC
CTGCCCGACTCCACCAGGCAGAAGATCGTGGAACTGGCGCACAGCGGCGCACGTCCCTGC
GACATTTCTCGGATTCTGCAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100931 exon2:KNOWN_protein_coding
GTGTCCAACGGCTGTGTGAGTAAGATCTTAGGGAGATATTACGAGACTGGATCGATCCGA
CCCAGAGCAATCGGTGGCAGCAAACCCAGAGTAGCCACCCCAGAAGTGGTTAGCAAGATA
GCCCAGTATAAAAGAGAGTGCCCTTCCATCTTTGCATGGGAAATCCGAGACAGGTTGCTA
TCTGAGGGAGTCTGTACCAACGACAATATCCCCAGT
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100932 exon3:KNOWN_protein_coding
GTGTCATCAATAAACCGAGTGCTGCGCAACCTGGCGAGCGAAAAGCAACAGATGGGCGCC
GATGGCATGTACGACAAGCTCAGGATGCTGAATGGGCAAACTGGGACCTGGGGGACCCGG
CCAGGGTGGTACCCCGGCACCTCGGTACCTGGCCAGCCAGCACAGG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100941 exon4:KNOWN_protein_coding
ACGGGTGTCAGCCGCAAGAAGGAGGAGGAGGAGGAGAAAACACAAACTCAATCAGCTCCA
ATGGCGAAGACTCAGACGAGGCCCAAATGAGGCTTCAGCTGAAGAGAAAATTACAAAGGA
ACAGAACATCTTTTACCCAGGAACAAATAGAGGCCCTAGAAAAAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100934 exon5:KNOWN_protein_coding
AATTTGAACGAACACATTACCCCGACGTGTTTGCCAGGGAAAGATTAGCTGCCAAAATCG
ACCTGCCAGAAGCAAGAATACAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100935 exon6:KNOWN_protein_coding
GTATGGTTCTCCAACAGAAGAGCAAAATGGAGAAGGGAGGAAAAACTTCGAAACCAGAGA
AGGCAGGCCAGTAACACACCCAGCCACATTCCCATTAGCAGTAGTTTCAGTACGAGCGTC
TACCAGCCAATCCCACAGCCTACCACACCAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100942 exon7:KNOWN_protein_coding
TGTCCTCTTTCACATCGGGTTCCATGCTGGGCAGAACGGACACAGCATTGACAAACTCCT
ACAGTGCGCTGCCACCTATGCCTAGTTTTACAATGGGCAACAACCTACCTATGCAA
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000316156 exon8:KNOWN_protein_coding
CCCCCCCCCCCCCCCACACACACACACACCTATCTTTTCCTGAGTTCCAATG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000408902 exon9:KNOWN_protein_coding
CAATGTGCCCAAACACTACAACGTATGATCCTTATGGACCCTTTATAAGGAACCCTAGGC
ATAGGCATGGAAACTGTCAGCCACAAAGTTCCAAAGGGACAAACCTAAAAT
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100943 exon10:KNOWN_protein_coding
GTCTCATTTCCCCTGGAGTGTCAGTCCCAGTTCAAGTACCCGGCAGTGAACCTGACATGT
CTCAGTACTGGCCAAGACTACAGTAA
I use Primer3Plus. The only settings I change is the product size range and the GC content. I want a product size that is ideally between 600 - 1000 base pairs. Under 400 is too short.
If you want to see how many nucleotides are in your sequence, you can go to LetterCounter.net and paste the text in there. This should give you an idea of what your product size will be.
Now that I have the exons sequence from 2 exons, there are several places I can go to generate primers. I want to have one primer on Exon 4 and the second on Exon 7. I copy and paste this sequence into Primer3Plus.
Primer Set 1 - Exon 4/5/6/7
Product Size - 531 bp
The first (forward) primer is on Exon 4, as I wanted. I can see from the second (reverse primer in yellow) may not be on Exon 7, but Exon 6.
While Primer3Plus will highlight the sequence for you, in the box below with the Pair the sequence will be reversed in order and reverse complimented. For instance, in the picture you can see Right Primer 3 is GAACCCGATGTGAAAGAGGA, even though the highlighted sequence is TCCTCTTTC ACATCGGGTT C.
In order to double check I will need to reverse compliment the sequence and search in my Ensembl text file to see what Exon its on. You can maybe do this in your head, but what I do is list the nucleodtides and work backwards. First I list the reverse compliment to the nucleotides. Then I reverse the whole order.
1) TCCTCTTTC ACATCGGGTT C (original primer seqeunce)
2) AGGAGAAAG TGTAGCCCAA G (reverse compliment to original)
3) G AACCCGATGT GAAAGAGGA (flipped sequence order)
Next, I take this sequence and search in the Ensembl text.
1) TCCTCTTTC ACATCGGGTT C
I do a search for ACATCGGG and I find that the primer is indeed on Exon 7.
Next I make another set with a primer on Exon 5 and Exon 10. This will give me a total of 4 primers, that I can use to mix and match (should one of the primers prove to be a poor choice).
Primer Set 2 - Exon 5/6/7/8/9/10
Next, I make a spreadsheet for the primers to keep track of what I order.
Next, I want to check to see what my PCR product should be. I enter in the sequence and my forward and reverse primer into a PCR Test, which is online at http://www.bioinformatics.org.
The results tell me the product size should be 516 bp, well within my desired range.
Now that I have the primer sets designed, I know the final product size is optimal, and that the GC content is above at least 45%, I can order them from a company. We use IDT, Integrated DNA technologies to order our primers.
From the IDT main ordering menu, I chose the Custom Synthesis -> Custom DNA oligos. On the order page, I enter in the sequences. All the default settings are fine.
If you are looking de novo for orthologue (gene equivalents) in another species, you may have to do some BLASTS to try to find them, including BLASTs for proteins, mRNA or highly conserved regions (like a promoter), depending on the amount of time diverged.
To begin, you have to search for the genes you want and save the sequence to a file. If you use Ensembl, you can search for a gene in a species and use the gene browser to visualize the gene structure. For example, I searched for Pax6 in the frog Xenopus. I can see right away that there are two isoforms of this gene.
So I know that the gene is spliced alternatively in two forms, which may have tissue specificity or functional importance. One isoform may be predominantly expressed, while another is found in low levels. Ideally, I want to capture them both. I will choose exons that are common to them both. For Pax6, the last 2 exons appear to be similiar enough. I want to export the exon sequence for two exons from Ensembl.
I want my oligo to span 2 exons ideally, such that the sequence spans an intron on either side. After each > is an exon. I find the 2 exons I want to use and copy/paste them into Primer3Plus.
I paste the exons into a text file and begin looking for a a good stretch, that spans introns, has a good GC content, and will give an appropriate product size.
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100940 exon1:KNOWN_protein_coding
ATGTCCCTAGGTCACAGCGGAGTCAATCAACTCGGGGGAGTGTTTGTGAACGGCCGACCC
CTGCCCGACTCCACCAGGCAGAAGATCGTGGAACTGGCGCACAGCGGCGCACGTCCCTGC
GACATTTCTCGGATTCTGCAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100931 exon2:KNOWN_protein_coding
GTGTCCAACGGCTGTGTGAGTAAGATCTTAGGGAGATATTACGAGACTGGATCGATCCGA
CCCAGAGCAATCGGTGGCAGCAAACCCAGAGTAGCCACCCCAGAAGTGGTTAGCAAGATA
GCCCAGTATAAAAGAGAGTGCCCTTCCATCTTTGCATGGGAAATCCGAGACAGGTTGCTA
TCTGAGGGAGTCTGTACCAACGACAATATCCCCAGT
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100932 exon3:KNOWN_protein_coding
GTGTCATCAATAAACCGAGTGCTGCGCAACCTGGCGAGCGAAAAGCAACAGATGGGCGCC
GATGGCATGTACGACAAGCTCAGGATGCTGAATGGGCAAACTGGGACCTGGGGGACCCGG
CCAGGGTGGTACCCCGGCACCTCGGTACCTGGCCAGCCAGCACAGG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100941 exon4:KNOWN_protein_coding
ACGGGTGTCAGCCGCAAGAAGGAGGAGGAGGAGGAGAAAACACAAACTCAATCAGCTCCA
ATGGCGAAGACTCAGACGAGGCCCAAATGAGGCTTCAGCTGAAGAGAAAATTACAAAGGA
ACAGAACATCTTTTACCCAGGAACAAATAGAGGCCCTAGAAAAAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100934 exon5:KNOWN_protein_coding
AATTTGAACGAACACATTACCCCGACGTGTTTGCCAGGGAAAGATTAGCTGCCAAAATCG
ACCTGCCAGAAGCAAGAATACAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100935 exon6:KNOWN_protein_coding
GTATGGTTCTCCAACAGAAGAGCAAAATGGAGAAGGGAGGAAAAACTTCGAAACCAGAGA
AGGCAGGCCAGTAACACACCCAGCCACATTCCCATTAGCAGTAGTTTCAGTACGAGCGTC
TACCAGCCAATCCCACAGCCTACCACACCAG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100942 exon7:KNOWN_protein_coding
TGTCCTCTTTCACATCGGGTTCCATGCTGGGCAGAACGGACACAGCATTGACAAACTCCT
ACAGTGCGCTGCCACCTATGCCTAGTTTTACAATGGGCAACAACCTACCTATGCAA
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000316156 exon8:KNOWN_protein_coding
CCCCCCCCCCCCCCCACACACACACACACCTATCTTTTCCTGAGTTCCAATG
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000408902 exon9:KNOWN_protein_coding
CAATGTGCCCAAACACTACAACGTATGATCCTTATGGACCCTTTATAAGGAACCCTAGGC
ATAGGCATGGAAACTGTCAGCCACAAAGTTCCAAAGGGACAAACCTAAAAT
>ENSXETG00000008175:ENSXETT00000017931 ENSXETE00000100943 exon10:KNOWN_protein_coding
GTCTCATTTCCCCTGGAGTGTCAGTCCCAGTTCAAGTACCCGGCAGTGAACCTGACATGT
CTCAGTACTGGCCAAGACTACAGTAA
I use Primer3Plus. The only settings I change is the product size range and the GC content. I want a product size that is ideally between 600 - 1000 base pairs. Under 400 is too short.
If you want to see how many nucleotides are in your sequence, you can go to LetterCounter.net and paste the text in there. This should give you an idea of what your product size will be.
Now that I have the exons sequence from 2 exons, there are several places I can go to generate primers. I want to have one primer on Exon 4 and the second on Exon 7. I copy and paste this sequence into Primer3Plus.
Primer Set 1 - Exon 4/5/6/7
Product Size - 531 bp
ACGGGTGTCAGCCGCAAGAAGGAGGAGGAGGAGGAGAAAACACAAACTCAATCAGCTC
CAATGGCGAAGACTCAGACGAGGCCCAAATGAGGCTTCAGCTGAAGAGAAAATTACAAA
GGAACAGAACATCTTTTACCCAGGAACAAATAGAGGCCCTAGAAAAAGAATTTGAACGAA
CACATTACCCCGACGTGTTTGCCAGGGAAAGATTAGCTGCCAAAATCGACCTGCCAGAAG
CAAGAATACAGGTATGGTTCTCCAACAGAAGAGCAAAATGGAGAAGGGAGGAAAAACTT
CGAAACCAGAGAAGGCAGGCCAGTAACACACCCAGCCACATTCCCATTAGCAGTAGTTTC
AGTACGAGCGTCTACCAGCCAATCCCACAGCCTACCACACCAGTGTCCTCTTTCACATCG
GGTTCCATGCTGGGCAGAACGGACACAGCATTGACAAACTCCTACAGTGCGCTGCCACC
TATGCCTAGTTTTACAATGGGCAACAACCTACCTATGCAAThe first (forward) primer is on Exon 4, as I wanted. I can see from the second (reverse primer in yellow) may not be on Exon 7, but Exon 6.
While Primer3Plus will highlight the sequence for you, in the box below with the Pair the sequence will be reversed in order and reverse complimented. For instance, in the picture you can see Right Primer 3 is GAACCCGATGTGAAAGAGGA, even though the highlighted sequence is TCCTCTTTC ACATCGGGTT C.
In order to double check I will need to reverse compliment the sequence and search in my Ensembl text file to see what Exon its on. You can maybe do this in your head, but what I do is list the nucleodtides and work backwards. First I list the reverse compliment to the nucleotides. Then I reverse the whole order.
1) TCCTCTTTC ACATCGGGTT C (original primer seqeunce)
2) AGGAGAAAG TGTAGCCCAA G (reverse compliment to original)
3) G AACCCGATGT GAAAGAGGA (flipped sequence order)
Next, I take this sequence and search in the Ensembl text.
1) TCCTCTTTC ACATCGGGTT C
I do a search for ACATCGGG and I find that the primer is indeed on Exon 7.
Next I make another set with a primer on Exon 5 and Exon 10. This will give me a total of 4 primers, that I can use to mix and match (should one of the primers prove to be a poor choice).
Primer Set 2 - Exon 5/6/7/8/9/10
Product Size - 549
AATTTGAACGAACACATTACCCCGACGTGTTTGCCAGGGAAAGATTAGCTGCCAAAATCGA
AATTTGAACGAACACATTACCCCGACGTGTTTGCCAGGGAAAGATTAGCTGCCAAAATCGA
CCTGCCAGAAGCAAGAATACAGGTATGGTTCTCCAACAGAAGAGCAAAATGGAGAAGGGA
GGAAAAACTTCGAAACCAGAGAAGGCAGGCCAGTAACACACCCAGCCACATTCCCATTAG
CAGTAGTTTCAGTACGAGCGTCTACCAGCCAATCCCACAGCCTACCACACCAGTGTCCTCT
TTCACATCGGGTTCCATGCTGGGCAGAACGGACACAGCATTGACAAACTCCTACAGTGCG
CTGCCACCTATGCCTAGTTTTACAATGGGCAACAACCTACCTATGCAACCCCCCCCCCCCC
CCACACACACACACACCTATCTTTTCCTGAGTTCCAATGCAATGTGCCCAAACACTACAA
CTATGATCCTTATGGACCCTTTATAAGGAACCCTAGGCATAGGCATGGAAACTGTCAGCCA
CAAAGTTCCAAAGGGACAAACCTAAAATGTCTCATTTCCCCTGGAGTGTCAGTCCCAGTT
CAAGTACCCGGCAGTGAACCTGACATGTCTCAGTACTGGCCAAGACTACAGTAANext, I make a spreadsheet for the primers to keep track of what I order.
Next, I want to check to see what my PCR product should be. I enter in the sequence and my forward and reverse primer into a PCR Test, which is online at http://www.bioinformatics.org.
The results tell me the product size should be 516 bp, well within my desired range.
Now that I have the primer sets designed, I know the final product size is optimal, and that the GC content is above at least 45%, I can order them from a company. We use IDT, Integrated DNA technologies to order our primers.
From the IDT main ordering menu, I chose the Custom Synthesis -> Custom DNA oligos. On the order page, I enter in the sequences. All the default settings are fine.
Subscribe to:
Posts (Atom)









.jpg)